Automated DNA Sequencers
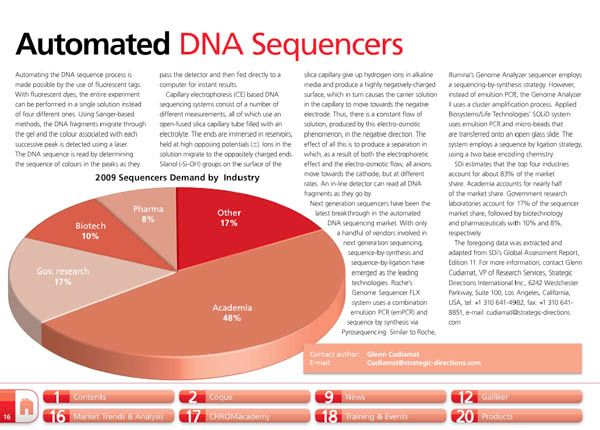
Automating the DNA sequence process is made possible by the use of fluorescent tags. With fluorescent dyes, the entire experiment can be performed in a single solution instead of four different ones. Using Sanger-based methods, the DNA fragments migrate through the gel and the colour associated with each successive peak is detected using a laser. The DNA sequence is read by determining the sequence of colours in the peaks as they pass the detector and then fed directly to a computer for instant results.
Automating the DNA sequence process is made possible by the use of fluorescent tags. With fluorescent dyes, the entire experiment can be performed in a single solution instead of four different ones. Using Sanger-based methods, the DNA fragments migrate through the gel and the colour associated with each successive peak is detected using a laser. The DNA sequence is read by determining the sequence of colours in the peaks as they pass the detector and then fed directly to a computer for instant results.

Fundamentals of Benchtop GC–MS Data Analysis and Terminology
April 5th 2025In this installment, we will review the fundamental terminology and data analysis principles in benchtop GC–MS. We will compare the three modes of analysis—full scan, extracted ion chromatograms, and selected ion monitoring—and see how each is used for quantitative and quantitative analysis.
Characterizing Plant Polysaccharides Using Size-Exclusion Chromatography
April 4th 2025With green chemistry becoming more standardized, Leena Pitkänen of Aalto University analyzed how useful size-exclusion chromatography (SEC) and asymmetric flow field-flow fractionation (AF4) could be in characterizing plant polysaccharides.
This information is supplementary to the article “Accelerating Monoclonal Antibody Quality Control: The Role of LC–MS in Upstream Bioprocessing”, which was published in the May 2025 issue of Current Trends in Mass Spectrometry.