Rethinking Chromatography Workflows with AI and Machine Learning
Excitement about using artificial intelligence (AI) and machine learning (ML) to accelerate drug development and identify better drug candidates is higher than ever. What about applying AI/ML to separation science?
Chromatographic techniques such as size-exclusion chromatography (SEC), capillary electrophoresis SDS (CE-SDS), and hydrophobic interaction chromatography (HIC) are integral for assessing the developability of biotherapeutics such as monoclonal or multispecific antibodies. Published research on AI and ML chromatography applications range from predicting retention times (1) to automating peak integration (2). In our own conversations with developability and protein production groups, we’ve heard interest in using historical data to predict molecular properties of unknown samples. We’ve also discussed using AI to predict optimal separation conditions, developing algorithms to flag compounds with poor chromatographic profiles, or even correlating characterization properties such as aggregation or glycosylation with molecular sequence—eventually making predictions on how to improve the sequences.
But before using AI/ML to analyze chromatographic data (or any other mode of data), you must provide both high-quality training data and well-annotated relevant metadata. This, in turn, hinges on having the appropriate workflows in place for the collection and management of chromatography data. We recently spoke with developability and protein production groups at major biopharma organizations across the industry about their data workflows and discovered several shared challenges that impact not only the ability to use AI/ML, but overall operational efficiency and effectiveness.
This article explores the key hurdles that must be overcome before we can use AI/ML with separation science data. We then offer an approach to managing these data that can overcome these challenges.
Analysis Not Comparable
Chromatography systems used during biotherapeutic development for analytical assessment and purification vary greatly depending on the application at hand. This includes the desired scale, speed, resolution, properties of the molecule being purified, and compliance requirements. Moreover, these instruments are made by several different, major vendors, each of whom offer their own software for primary data analysis (peak detection and quantification). Analysis practices may also differ between scientists and groups. This may be fine for a single experiment, but it means that results from multiple experiments cannot be easily or validly compared. When it comes to creating training data for AI algorithms, inconsistency in analysis and resulting noise or error could impact model performance (3).
Disjointed Files and Scattered Metadata
As with many other types of data, scientists working with chromatography data must assess or reassess results in the context of additional metadata not stored in instrument software, or even new assay data. For example, they need additional information on control samples and molecular properties (Ɛ, pI, A280, molecular weight) to calculate and interpret key results (monomer or multimer %, retention time). They may compare purification efficiency across different buffer conditions or judge the results against other, non-chromatographic assays, such as biophysical assays. For such secondary analyses, static visualizations, such as .png or .jpg screen captures, decentralized storage of metadata in disjointed files, and spreadsheet-heavy workflows lead to a lack of both traceability and scalability. Ensuring proper contextualization of results with key metadata is also important for AI and ML, for which accurate annotations are important.
Manual Reporting to Disparate Locations
Different teams or scientists within the same group must often manually consolidate their results alongside the chromatograms so that they can be reported to an electronic lab notebook (ELN) or a central data store. Due to the time and effort needed for this, results might not even be registered in the first place. This hinders collaboration and the ability to learn from previous projects. This includes future learning by AI and ML, which requires that training datasets be complete and well-balanced; if data are selectively or inconsistently reported, the chances of generating such datasets is lowered.
Challenges in chromatography data workflows: research and development groups analyze data on different instruments, making it hard to compare results; secondary analysis using metadata stored in disjointed files; and manual reporting to disparate locations. Click the figure to enlarge.
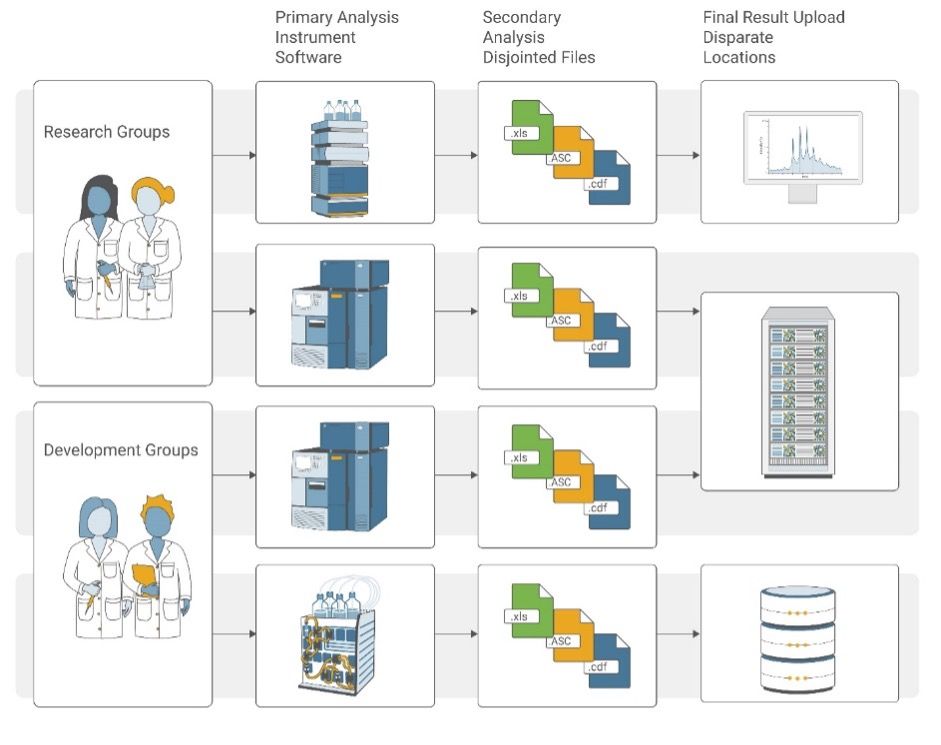
Challenges in chromatography data workflows: research and development groups analyze data on different instruments, making it hard to compare results; secondary analysis using metadata stored in disjointed files; and manual reporting to disparate locations.
Rethinking Chromatography Data Systems
Considering these challenges, we believe the industry needs to look towards building more centralized, integrated software systems for chromatographic data. Key features of these systems must include vendor-agnostic integrations with industry-standard instruments that will bring together data from various applications into a single analysis platform, for analytical assessment and purification of molecules. Key metadata about molecular properties, production cell lines, and experimental conditions, as well as data from non-chromatographic assays, could be automatically retrieved through integration with parallel analytical or registration and tracking systems. The system should offer automated calculation, display, filtering, and ranking of samples, with the ability to customize methods at an organizational level, using APIs.Reporting of results should also be simplified, through close integration with surrounding IT infrastructure and registration systems. This ensures proper data governance and makes data exchange, joint decision-making, and use of historical data easier.
A single platform for all chromatography analysis, from metadata import to result calculation, with streamlined, automated reporting to surrounding systems. Click the figure to enlarge.

Community-built solutions are one option towards building this type of system. In contrast, commercially built solutions from firms combining expertise in software development and biopharmaceutical research may better serve enterprise needs surrounding scalability and compliance requirements, with stable maintenance and support and a long-term product roadmap. Commercial systems can still be open, providing frameworks to implement novel algorithms, for example ML algorithms for quality assessment of complex molecules.
Eventually, with the benefit of centralization and reliable annotation, such a software system has the potential to achieve many of the AI-based objectives we have heard about the system might learn from scientist input so that it can automate quality control, or it might use sequence metadata to predict sequences that produce molecules with more desirable characteristics. With a concrete plan and the right solutions in place for data analysis and management, separation scientists will then be positioned to leverage AI/ML for faster, more successful drug development and higher ROI.
References
- Singh, Y. R.; Shah, D. B.; Maheshwari, D. G.; Shah, J. S.; Shah, S. Advances in AI-Driven Retention Prediction for Different Chromatographic Techniques: Unraveling the Complexity. Critical Reviews in Analytical Chemistry 2024.
- Satwekar, A.; Panda, A.; Nandula, P.; Sripada, S.; Govindaraj, R.; Rossi, M. Digital by Design Approach to Develop a Universal Deep Learning AI Architecture for Automatic Chromatographic Peak Integration. Biotechnology and Bioengineering 2023, 120 (7), 1822–1843. https://doi.org/10.1002/bit.28406.
- Budach, L.; Feuerpfeil, M.; Ihde, N.; Nathansen, A.; Noack, N.; Patzlaff, H.; Naumann, F.; Harmouch, H. The Effects of Data Quality on Machine Learning Performance. arXiv November 9, 2022. https://doi.org/10.48550/arXiv.2207.14529.
How Many Repetitions Do I Need? Caught Between Sound Statistics and Chromatographic Practice
April 7th 2025In chromatographic analysis, the number of repeated measurements is often limited due to time, cost, and sample availability constraints. It is therefore not uncommon for chromatographers to do a single measurement.
Fundamentals of Benchtop GC–MS Data Analysis and Terminology
April 5th 2025In this installment, we will review the fundamental terminology and data analysis principles in benchtop GC–MS. We will compare the three modes of analysis—full scan, extracted ion chromatograms, and selected ion monitoring—and see how each is used for quantitative and quantitative analysis.
Characterizing Plant Polysaccharides Using Size-Exclusion Chromatography
April 4th 2025With green chemistry becoming more standardized, Leena Pitkänen of Aalto University analyzed how useful size-exclusion chromatography (SEC) and asymmetric flow field-flow fractionation (AF4) could be in characterizing plant polysaccharides.

.png&w=3840&q=75)

.png&w=3840&q=75)



.png&w=3840&q=75)



.png&w=3840&q=75)
